Use the Pandas Dataframe Deptstats Again
Getting Started
How to Avoid a Pandas Pandemonium
A deep dive into common Pandas mistakes. Role I: Writing skilful code and spotting silent failures.
When you lot showtime start out using Pandas, it's often best to just get your anxiety moisture and deal with problems equally they come up. And so, the years pass, the astonishing things you've been able to build with it start to accumulate, only you have a vague inkling that you lot keep making the same kinds of mistakes and that your code is running really slowly for what seems like pretty unproblematic operations. This is when it's time to dig into the inner workings of Pandas and have your lawmaking to the next level. Like with any library, the best style to optimize your code is to empathise what's going on underneath the syntax.
Start in Part I, nosotros're going to eat our vegetables and cover writing clean code and spotting common silent failures. So in Part II, nosotros'll get to speeding up your runtime and lowering your memory footprint.
I besides made a Jupyter notebook with the whole lesson, both parts included.
First, permit's make some fake data to play with. I'll make a large DataFrame to illustrate larger processing problems and a small-scale DataFrame to illustrate point changes in a localized fashion.
# for neatness, it helps to keep all of your imports up top
import sys
import traceback
import numba
import numpy as np
import pandas as pd
import numpy.random as nr
import matplotlib.pyplot equally plt% matplotlib inline
# generate some fake data to play with
data = {
"day_of_week": ["Monday", "Tuesday", "Wednesday", "Thursday", "Friday", "Saturday", "Sunday"] * 1000,
"booleans": [True, Fake] * 3500,
"positive_ints": nr.randint(0, 100, size=7000),
"mixed_ints": nr.randint(-100, 100, size=7000),
"lat1": nr.randn(7000) * 30,
"lon1": nr.randn(7000) * 30,
"lat2": nr.randn(7000) * xxx,
"lon2": nr.randn(7000) * xxx,
}df_large = pd.DataFrame(information)
df_large.head()
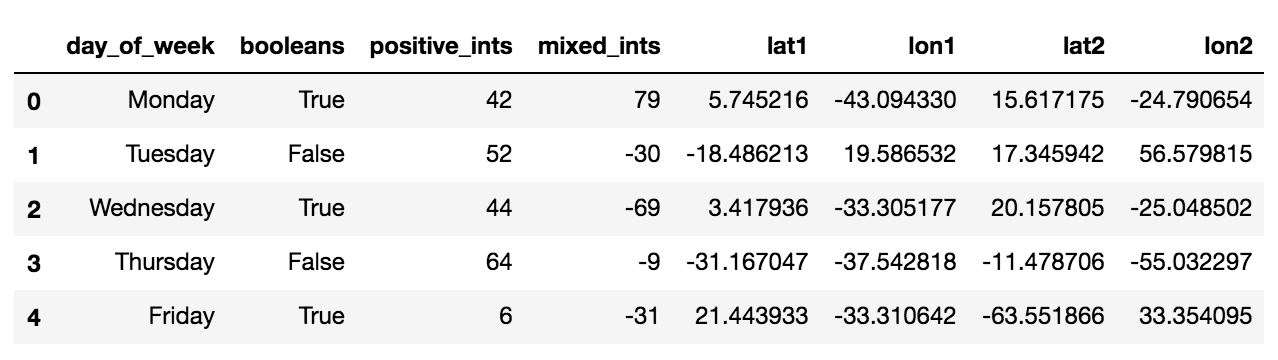
small = {
'a': [1, 1],
'b': [2, 2]
} df_small = pd.DataFrame(small) df_small

Great, at present nosotros're all gear up up!
Writing practiced code
Earlier we do the "cool" stuff like writing faster and more memory-optimized code, nosotros need to do it on a foundation of some fairly mundane-seeming coding best practices. These are the little things, such every bit naming things expressively and writing sanity checks, that volition help continue your code maintainable and readable by your peers.
Sanity checking is elementary and totally worth it
Just because this is mostly simply data assay, and it might non make sense to put up a whole suite of unit tests for it, doesn't hateful you can't exercise any kind of checks at all. Peppering your notebook lawmaking with assert tin get a long way without much actress piece of work.
In a higher place, we fabricated a DataFrame df_large that contains numbers with some pre-divers rules. For example, you lot can check for data entry errors by trimming whitespace and checking that the number of entries stays the aforementioned:
large_copy = df_large.copy() assert large_copy["day_of_week"].str.strip().unique().size == large_copy["day_of_week"].unique().size
Naught should happen if y'all run this lawmaking. However, modify information technology slightly to break it and…
large_copy.loc[0, "day_of_week"] = "Mon " affirm large_copy["day_of_week"].str.strip().unique().size == large_copy["day_of_week"].unique().size
Bam! AssertionError. Suppose you had some code that drops duplicates and ships the delta to a client. Trivial whitespace additions would be missed and sent off as its own unique information point. Information technology'due south not a big deal when it's the days of the week and you tin spot check it easily, just if information technology's potentially thousands or more than of unique strings, these kinds of checks tin can save yous a large headache.
Use consequent indexing
Pandas grants you a lot of flexibility in indexing, but information technology tin add upwards to a lot of confusion later on if you're not disciplined virtually keeping a consistent style. This is one proposed standard:
# for getting columns, use a string or a list of strings for multiple columns
# note: a one-column DataFrame and a Serial are not the same thing
one_column_series = df_large["mixed_ints"]
two_column_df = df_large[["mixed_ints", "positive_ints"]] # for getting a 2D slice of data, use `loc`
data_view_series = df_large.loc[:10, "positive_ints"]
data_view_df = df_large.loc[:10, ["mixed_ints", "positive_ints"]] # for getting a subset of rows, as well use `loc`
row_subset_df = df_large.loc[:10, :]
This is just a combination of what I personally use and what I've seen others practice. Here's another set of ways to do the exact same things every bit above, but yous may come across why they either don't have equally much adoption or are discouraged from use.
# one style is to use `df.loc` for everything, only information technology tin can look clunky
one_column_series = df_large.loc[:, "mixed_ints"]
two_column_df = df_large.loc[:, ["mixed_ints", "positive_ints"]] # you can use `iloc`, which is `loc` just with indexes, only it's not as clear
# also, you lot're in problem if you ever change the column order
data_view_series = df_large.iloc[:10, two]
data_view_df = df_large.iloc[:10, [3, two]] # yous tin go rows similar you slice a listing, merely this can be confusing
row_subset_df = df_large[:x]
Here's a little gotcha about that final example: df_large[:10] gets you the first x rows, but df_large[10] gets you lot the tenth column. This is why it'southward so important to index things every bit clearly as possible, even if it means you have to do more typing.
But don't use chained indexing
What is chained indexing? It's when you separately index the columns and the rows, which will make 2 separate calls to __getitem__() (or worse, one phone call to __getitem__() and one to __setitem__() if y'all're making assignments, which we demonstrate below). It's not and so bad if you're but indexing and not making assignments, but information technology'south nevertheless not ideal from a readability standpoint because if yous index the rows in i place and then index a column, unless you lot're very disciplined about variable naming, information technology'southward easy to lose runway of what exactly y'all indexed.
# this is what chained indexing looks like
data_view_series = df_large[:x]["mixed_ints"]
data_view_df = df_large[:10][["mixed_ints", "positive_ints"]] # this is likewise chained indexing, only low-key
row_subset_df = df_large[:x]
data_view_df = row_subset_df[["mixed_ints", "positive_ints"]]
The 2nd case isn't something people usually intend to exercise, it often happens when people alter DataFrames in nested for loops, where first y'all iterate through each row, but and so you want to do something to some of the columns. If you get a SettingWithCopyWarning, try to await for these kinds of patterns or anything where you might be separately slicing rows and and then slicing columns (or vice versa) and replace them with df.loc[rows, columns].
Common silent failures
Fifty-fifty if you do all of the above, sometimes Pandas' flexibility can lull you into making mistakes that don't actually make yous mistake out. These are particularly pernicious considering yous often don't realize something is wrong until something far downstream doesn't make sense, and it's very difficult to trace back to what the cause was.
View vs. Copy
A view and a copy of a DataFrame can look identical to you in terms of the values it contains, merely a view references a piece of an existing DataFrame and a copy is a whole unlike DataFrame. If you change a view, you change the existing DataFrame, just if you modify a copy, the original DataFrame is unaffected. Make sure you aren't modifying a view when you think you're modifying a re-create and vice versa.
It turns out that whether you're dealing with a copy or a view is very difficult to predict! Internally, Pandas tries to optimize by returning a view or a re-create depending on the DataFrame and the actions yous take. Yous tin can force Pandas to make a copy for you by using df.copy() and you can force Pandas to operate in place on a DataFrame by setting inplace=Truthful when it's available.
When to brand a copy and when to use a view? It'south difficult to say for sure, only if your data is small or your resources are large and you desire to go functional and stateless, you tin try making a re-create for every performance, like Spark would, since it's probably the safest way to practice things. On the other hand, if you lot have lots of data and a regular laptop, y'all might desire to operate in place to prevent your notebooks from crashing.
# intentionally making a copy will ever make a copy
small_copy = df_small.copy() small_copy

While they wait identical, this is actually a totally separate data object than the df_small we made earlier.
Allow's await at what inplace=True does. df.drop() is method that allows yous to drop columns from a DataFrame, and information technology's one of the Pandas DataFrame methods that let y'all to specify if you want the performance to occur in-place or if you want information technology to return a new object. Returning a new object, or inplace=False is e'er the default.
small_copy.driblet("b", centrality=one) small_copy

What happened to the drop? Information technology returned a new object, but you didn't point to information technology with a variable, and then it disappeared. The original, small_copy, remains unmodified. Now let's plow on in-place modification:
small_copy.drop("b", axis=1, inplace=Truthful) small_copy

The original, small_copy, has now been inverse. Proficient thing nosotros made a re-create!
Permit's remake small_copy and make a betoken modification to see what I meant before well-nigh the original DataFrame existence a dissimilar object.
# let'south meet what happens if y'all assign to a copy
small_copy = df_small.copy() # yous should always use `loc` for consignment
small_copy.loc[0, 'b'] = 4 small_copy

# original is unchanged
df_small 
Watch out for out of order processing
In Jupyter notebooks, it'due south near unavoidable to change and re-process cells out of society — nosotros know nosotros shouldn't do it, simply it e'er ends up happening. This is a subset of the view vs. copy trouble because if you lot know that y'all're making a modify that fundamentally alters the properties of a column, you lot should consume the retentivity cost and make a new copy, or something like this might happen where you run the latter two cells in this block over and over and see the max value go pretty unstable.
large_copy = df_large.copy()
large_copy.loc[0, "positive_ints"] = 120
large_copy["positive_ints"].max() > 120
Retrieve, positive_ints was set to be between 0 and 100, meaning that setting the outset value to 120 means the max value has to exist 120 now. Simply what happens if I run a cell like this a few times?
large_copy["positive_ints"] = large_copy["positive_ints"] * 3
large_copy["positive_ints"].max() > 360
The first time you run it, information technology behaves as expected, but run it once more…
large_copy["positive_ints"] = large_copy["positive_ints"] * iii
large_copy["positive_ints"].max() > 1080
This example seems obvious, but it's really like shooting fish in a barrel to make cells that mutate data irreversibly and in a way that builds on itself every time yous re-run it. The best mode to avert this problem is to make a new DataFrame with a new variable name when you're starting a cell that volition make significant in-place data mutations.
large_copy_x3 = large_copy.copy()
large_copy_x3["positive_ints"] = large_copy["positive_ints"] * three
large_copy_x3["positive_ints"].max() > 360
No matter how many times you run that cake, it volition always render 360, simply every bit you originally expected.
Never ready errors to "ignore"
Some Pandas methods allow you to ignore errors by default. This is almost always a bad idea because ignoring errors means it just puts your unparsed input in place of where the output should have been. Let'due south make a Series with two normal dates and a Futurama-like one.
parsed_dates = pd.to_datetime(["ten/xi/2018", "01/xxx/1996", "04/15/9086"], format="%m/%d/%Y", errors="ignore") parsed_dates > array(['10/11/2018', '01/30/1996', '04/15/9086'], dtype=object)
Note in the example output that if y'all were non overly familiar with what Pandas outputs should exist, seeing an output of array type might not seem that unusual to yous, and you might just move on with your analysis, non knowing that something had gone wrong.
If you lot turn off errors="ignore", you will see the traceback:
Traceback (nigh recent call final):
File "<ipython-input-22-12145c38fd6e>", line 3, in <module>
pd.to_datetime(["10/11/2018", "01/thirty/1996", "04/15/9086"], format="%chiliad/%d/%Y")
pandas._libs.tslibs.np_datetime.OutOfBoundsDatetime: Out of bounds nanosecond timestamp: 9086-04-15 00:00:00 What happened here is that Python timestamps are indexed in nanoseconds, and that number is stored as an int64, so whatsoever year that is later than roughly 2262 volition overflow its ability to exist stored. As a information scientist, you may be forgiven for non knowing that little bit of esoterica, but this is just one of the many hidden idiosyncrasies buried in Python/Pandas, so ignore at your peril.
This is what a date Series is supposed to expect like, in case you lot wondering:
pd.to_datetime(["10/11/2018", "01/30/1996"], format="%m/%d/%Y") > DatetimeIndex(['2018-10-11', '1996-01-30'], dtype='datetime64[ns]', freq=None)
The object dtype can hide mixed types¶
Each Pandas column has a type, simply there is an uber-type called object, which means each value is actually just a pointer to some capricious object. This allows Pandas to have a neat deal of flexibility (i.eastward. columns of lists or dictionaries or whatsoever yous want!), merely it tin result in silent fails.
Spoiler alert: this won't exist the commencement time object type causes united states of america problems. I don't want to say you shouldn't utilize information technology, but once you're in product mode, yous should definitely use information technology with caution.
# we start out with integer types for our small data
small_copy = df_small.copy()
small_copy.dtypes >
a int64
b int64
dtype: object # reset ones of the column'south dtype to `object`
small_copy["b"] = small_copy["b"].astype("object")
small_copy.dtypes >
a int64
b object
dtype: object
Now allow's make some mischief again and mutate the b cavalcade with probably ane of those most frustrating silent failures out there.
small_copy["b"] = [4, "4"]
small_copy 
(I'g sure for a lot of you, this is the information equivalent of nails on a chalkboard.)
Now permit's say yous demand to drop duplicates and transport the event. If you dropped past column a, you would get something expected:
small_copy.drop_duplicates("a") 
But if you dropped past column b, this is what you would get:
small_copy.drop_duplicates("b") 
Tread advisedly with Pandas schema inference
When you lot load in a big, mixed-type CSV and Pandas gives you the option to set low_memory=Fake when it encounters some data it doesn't know how to handle, what information technology's actually doing is only making that entire column object type so that the numbers it can convert to int64 or float64 get converted, merely the stuff it can't convert just sit in that location every bit str. This makes the column values able to peacefully co-exist, for now. But one time you endeavour to do whatever operations on them, you'll encounter that Pandas was just trying to tell you all along that you lot can't presume all the values are numeric.
Note: Remember, in Python, NaN is a float! And then if your numeric column has them, cast them to float even if they're really int.
Let's create an intentionally birdbrained DataFrame and salve information technology somewhere and then nosotros can read information technology back in. This DataFrame has a combination of integers and a string that the (most likely human, probably working in Excel) writer thought was an indication of a NaN value just is actually not covered by the default NaN parsers.
mixed_df = pd.DataFrame({"mixed": [100] * 100 + ["-"] + [100] * 100, "ints": [100] * 201}) mixed_df.to_csv("test_load.csv", header=True, index=Imitation)
When you lot read it back in, Pandas will try to practice schema inference, but it will stumble on the mixed types column and non attempt to be too opinionated about it and leave the values every bit they are in an object cavalcade.
mixed_df = pd.read_csv("test_load.csv", header=0)
mixed_df.dtypes >
mixed object
ints int64
dtype: object
The all-time way to brand certain your data is interpreted correctly is to prepare your dtypes manually and specify your NaN characters. Information technology'southward a hurting, and you don't always have to do information technology, simply it'southward always a good idea to open the get-go few lines of whatsoever information file y'all take and make certain it tin can really exist automatically parsed.
mixed_df = pd.read_csv("test_load.csv", header=0, dtype={"mixed": bladder, "ints": int}, na_values=["-"])
mixed_df.dtypes >
mixed float64
ints int64
dtype: object
Decision, Function I
Pandas is probably the greatest matter since sliced lists, only it's machine-magic until it's not. To understand when and how you tin can let it take the cycle and when you have to do a manual override, you either demand to have a solid grasp of its internal workings, or you can be similar me and hitting your caput confronting them for half a decade and try to acquire from your mistakes. Every bit the saying goes, fool me once, shame on you. Fool me 725 times…tin can't fool me once more.
Please bank check out Part II: Speed-ups and memory optimizations.
Source: https://towardsdatascience.com/how-to-avoid-a-pandas-pandemonium-e1bed456530
0 Response to "Use the Pandas Dataframe Deptstats Again"
Post a Comment